Search
The discovery of CRISPR-Cas9 and its widespread use has revolutionised and propelled research in biological sciences.
We aimed to investigate the molecular basis underlying a novel phenotype including hypopituitarism associated with primary ovarian insufficiency.
Directed evolution emulates the process of natural selection to produce proteins with improved or altered functions. These approaches have proven to be very powerful but are technically challenging and particularly time and resource intensive. To bypass these limitations, we constructed a system to perform the entire process of directed evolution in silico.
Transcription of the human mitochondrial genome and correct processing of the two long polycistronic transcripts are crucial for oxidative phosphorylation. According to the tRNA punctuation model, nucleolytic processing of these large precursor transcripts occurs mainly through the excision of the tRNAs that flank most rRNAs and mRNAs.
Sarcomeric gene mutations are associated with the development of hypertrophic cardiomyopathy (HCM). Current drug therapeutics for HCM patients are effective in relieving symptoms, but do not prevent or reverse disease progression. Moreover, due to heterogeneity in the clinical manifestations of the disease, patients experience variable outcomes in response to therapeutics.
The evolutionary acquisition of mitochondria has given rise to the diversity of eukaryotic life. Mitochondria have retained their ancestral α-proteobacterial traits through the maintenance of double membranes and their own circular genome. Their genome varies in size from very large in plants to the smallest in animals and their parasites. The mitochondrial genome encodes essential genes for protein synthesis and has to coordinate its expression with the nuclear genome from which it sources most of the proteins required for mitochondrial biogenesis and function.
Changes in the rate and fidelity of mitochondrial protein synthesis impact the metabolic and physiological roles of mitochondria. Here we explored how environmental stress in the form of a high-fat diet modulates mitochondrial translation and affects lifespan in mutant mice with error-prone or hyper-accurate mitochondrial ribosomes. Intriguingly, although both mutations are metabolically beneficial in reducing body weight, decreasing circulating insulin and increasing glucose tolerance during a high-fat diet, they manifest divergent (either deleterious or beneficial) outcomes in a tissue-specific manner.
Transcriptomic technologies have revolutionized the study of gene expression and RNA biology. Different RNA sequencing methods enable the analyses of diverse species of transcripts, including their abundance, processing, stability, and other specific features. Mitochondrial transcriptomics has benefited from these technologies that have revealed the surprising complexity of its RNAs. Here we describe a method based upon cyclization of mitochondrial RNAs and next generation sequencing to analyze the steady-state levels and sizes of mitochondrial RNAs, their degradation products, as well as their processing intermediates by capturing both 5' and 3' ends of transcripts.
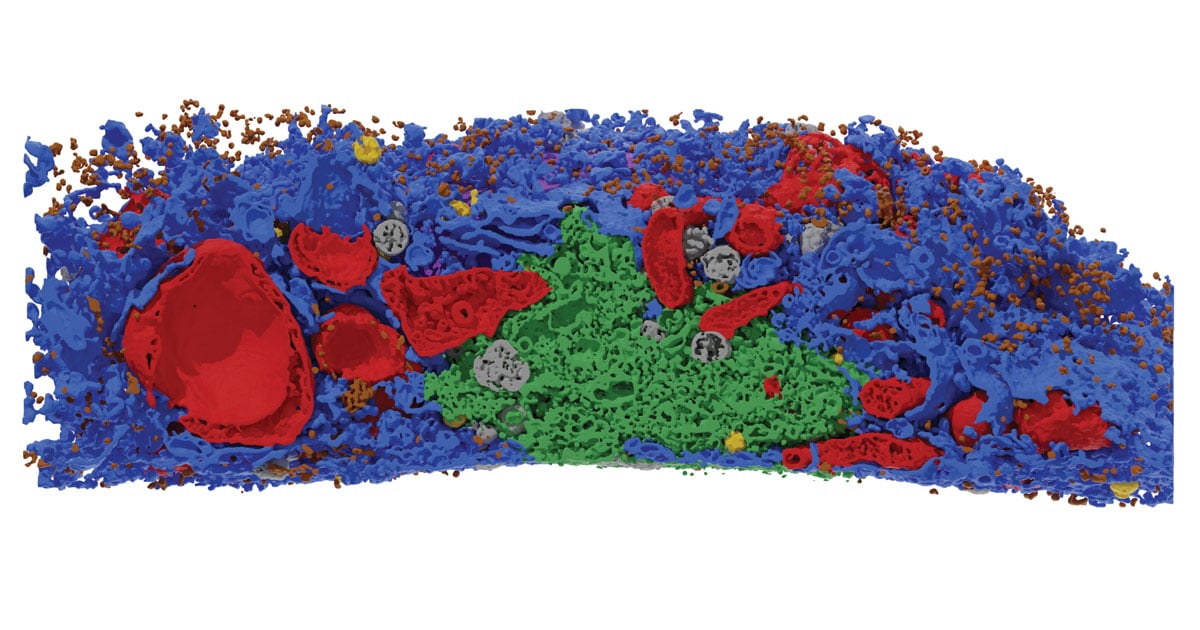
Researchers from The Kids Research Institute Australia and The University of Western Australia have developed a new technique to see inside cells with unprecedented detail, revealing a complicated web of interactions that provides new insights into how cells stay healthy.

The generous support of West Australians through Channel 7’s Telethon Trust will help support vital child health research at The Kids Research Institute Australia in 2023.
